ADAM
AMPs are potent drug candidates against microbial organisms such as bacteria, fungi, parasites, and viruses.
Several AMP databases were released in the past few years. Some databases are species-specific such as
BACTIBASE, BAGEL2, DADP, and PhytAMP; some are derived from a broad spectrum of species such as APD2, CAMP2,
and DAMPD. Among all of these databases, none contains all the AMP sequences and little is done for
AMP sequence-structure relationships in a large scale.
Lack of an easy access to link AMP sequences to structures and vice versa is a problem although AMPs are
known to have various structures. Additionally sequence-structure relationships are important, particularly
for antimicrobial drug design. To our knowledge, ADAM is the first AMP database to compile the AMP folds
comprehensively and establish associations between AMP sequences and structures systematically.
The size and sequence-structure analysis are two distant features of ADAM. ADAM comes from multiple sources.
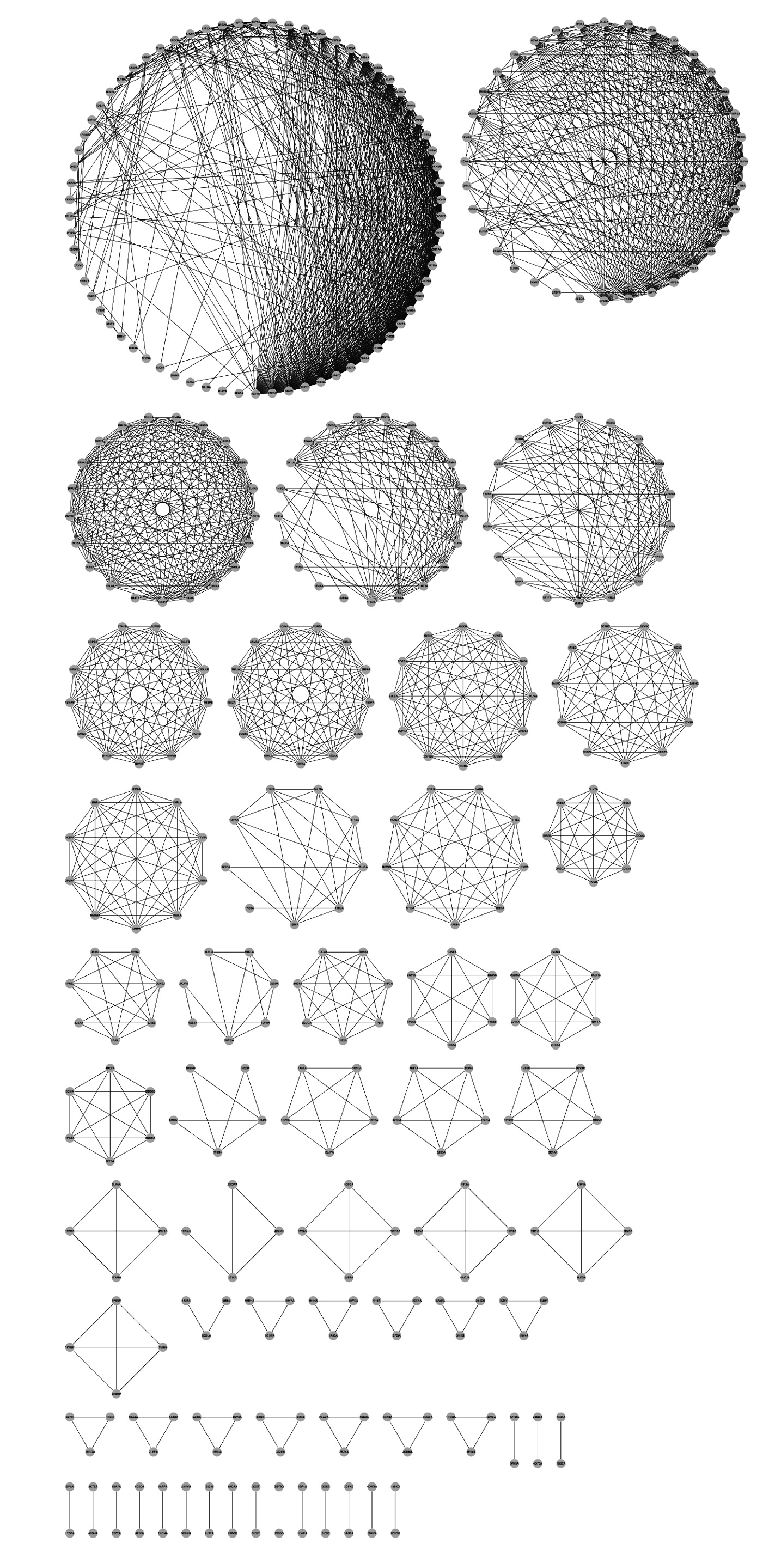
In the current release, ADAM is the largest AMP database, which contains 7,811 unique AMP sequences. To view the relationships between the AMP sequences and structures,
the best matching AMP structures.
The 136 AMP fold clusters annotated by CATH (Class, Architecture, Topology, and Homologous Superfamily),
SCOP(Class, Fold, Superfamily, Family), and Pfam (protein sequence conserved domain)would list the
associated AMP sequences or vice versa.
|
AMPs are potent drug candidates against microbial organisms such as bacteria, fungi, parasites, and viruses.
Several AMP databases were released in the past few years. Some databases are species-specific such as
BACTIBASE, BAGEL2, DADP, and PhytAMP; some are derived from a broad spectrum of species such as APD2, CAMP2,
and DAMPD. Among all of these databases, none contains all the AMP sequences and little is done for
AMP sequence-structure relationships in a large scale.
Lack of an easy access to link AMP sequences to structures and vice versa is a problem although AMPs are known to have various structures. Additionally sequence-structure relationships are important, particularly for antimicrobial drug design. To our knowledge, ADAM is the first AMP database to compile the AMP folds comprehensively and establish associations between AMP sequences and structures systematically. The size and sequence-structure analysis are two distant features of ADAM. ADAM comes from multiple sources. In the current release, ADAM is the largest AMP database, which contains 7,811 unique AMP sequences. To view the relationships between the AMP sequences and structures, the best matching AMP structures. The 136 AMP fold clusters annotated by CATH (Class, Architecture, Topology, and Homologous Superfamily), SCOP(Class, Fold, Superfamily, Family), and Pfam (protein sequence conserved domain)would list the associated AMP sequences or vice versa. |  |
